Research Briefing
Published: 01 September 2025
Nature Biotechnology
(2025)Cite this article
Subjects
Precision CRISPR–Cas9-mediated genome engineering remains challenging, particularly gene integration and editing in non-dividing cells. We present Pythia, a deep learning solution that forecasts optimal repair templates and enables predictable and accurate genome editing in diverse cellular contexts, both in vivo and in vitro.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
27,99 € / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
269,00 € per year
only 22,42 € per issue
Buy this article
Purchase on SpringerLink
Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Additional access options:
Log in
Learn about institutional subscriptions
Read our FAQs
Contact customer support
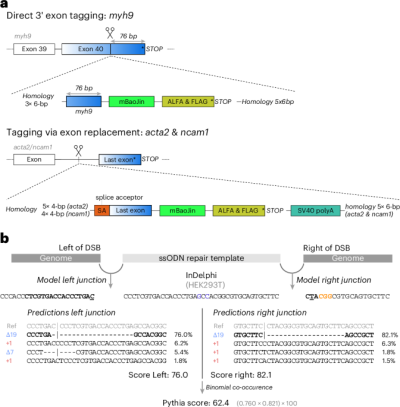
Fig. 1: Pythia provides AI-based predictions of repair template performance.
References Wang, J. Y. & Doudna, J. A. CRISPR technology: a decade of genome editing is only the beginning. Science 379, eadd8643 (2023). This review explores the rapid evolution of CRISPR genome editing over the past decade and outlines key advances, challenges and future directions across medicine, agriculture and basic research.
PubMed
CAS
Google Scholar
van Overbeek, M. et al. DNA repair profiling reveals nonrandom outcomes at Cas9-mediated breaks. Mol. Cell 63, 633–646 (2016). This study demonstrates that DNA repair outcomes following Cas9 cleavage are non-random and largely determined by the target sequence.
PubMed
Google Scholar
Shen, M. W. et al. Predictable and precise template-free CRISPR editing of pathogenic variants. Nature 563, 646–651 (2018). This study shows that template-free CRISPR–Cas9 editing can yield highly predictable repair outcomes at pathogenic loci, laying the groundwork for AI-driven models to anticipate editing results.
PubMed
PubMed Central
CAS
Google Scholar
Anzalone, A. V. et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576, 149–157 (2019). This study introduces prime editing, which uses a Cas9–reverse transcriptase and pegRNA-encoded template to install precise edits without double-strand breaks or separate donor DNA.
PubMed
PubMed Central
CAS
Google Scholar
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016). This study introduces cytosine base editing using a Cas9 nickase–cytidine deaminase fusion that converts C>T (G>A) without double-stranded breaks or donor DNA.
PubMed
PubMed Central
CAS
Google Scholar
Download references
Additional information Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This is a summary of: Naert, T. et al. Precise, predictable genome integrations by deep-learning-assisted design of microhomology-based templates. Nat. Biotechnol. https://doi.org/10.1038/s41587-025-02771-0 (2025).
Rights and permissions About this article
Cite this article Pythia provides deep learning-driven precision in CRISPR–Cas9 genome engineering.
Nat Biotechnol (2025). https://doi.org/10.1038/s41587-025-02818-2
Download citation
Published: 01 September 2025
DOI : https://doi.org/10.1038/s41587-025-02818-2
Associated content

Leave a Reply